Gene study provides AI insight into a leading cause of dementia
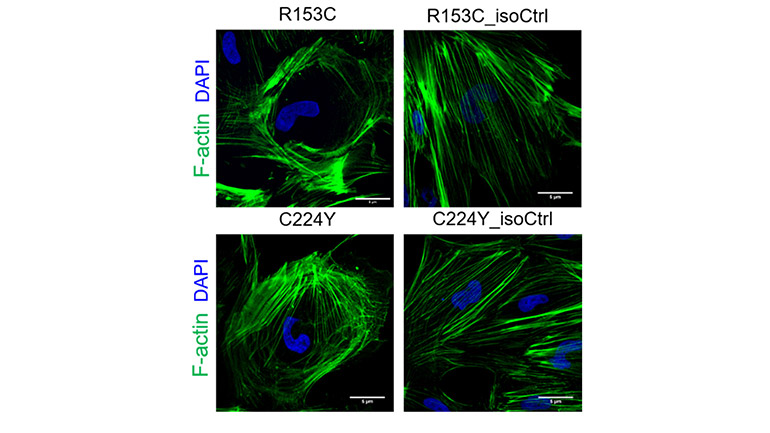
Immunofluorescence staining of F-actin in Induced Pluripotent Stem Cell-derived Vascular Smooth Muscle Cells derived from CADASIL and Isogenic control induced pluripotent stem cells.
New research has revealed a mechanism behind one of the leading genetic causes of vascular dementia and existing medication that could treat it, with the help of artificial intelligence expertise at the University of Leicester.
The new study, published in Science Advances, explores if posphodiesterase 5 (PDE5) inhibitors, an existing medication which breaks down cyclic guanosine monophosphate in cells, can reverse the functional abnormality and survival of mutant vascular smooth muscle cells (VSMCs).
Protecting vascular smooth muscle cells (VSMCs) is critical in cardiovascular medicine because they are the primary cells responsible for maintaining the structural integrity, elasticity, and contractility of the arterial wall. In the context of diseases like atherosclerosis, maintaining healthy, contractile VSMCs prevents the formation of vulnerable, rupture-prone plaques, thereby reducing the risk of heart attacks and strokes. They play a pivotal role in the initiation, progression, and stabilization of small vessel disease (SVD), acting as key moderators of vascular remodelling and dysfunction.
If left untreated, small vessel disease can cause repeated small strokes that gradually damage brain function, leading to problems with memory and thinking, and eventually dementia.
The research team, led by the University of Manchester and supported by the University of Leicester, focused their attention on a human gene called NOTCH3. This gene encodes a transmembrane receptor that is important for the differentiation and maturation of VSMCs.
Variants in NOTCH3 cause a condition known as cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL), a genetic small vessel disease that leads to stroke, dementia and premature death. Understanding how NOTCH3 variants result in clinical symptoms will help to produce effective drugs to treat small vessel disease and vascular dementia.
This project used human induced pluripotent stem cell (iPSC) models to develop ethical, patient-specific and an unlimited supply of human cells for disease modelling, drug discovery, and safety testing instead of using animal models. iPSCs are patient-derived, reprogrammed somatic cells (e.g. skin or blood) capable of differentiating into any cell type.
The scientists used these stem cell models to generate the brain-specific VSMCs that are vulnerable to NOTCH3 variants, with extensive extracellular matrix accumulation, impaired cell adhesion and nitric oxide signalling. They found that inhibiting PDE5 successfully reversed the functional abnormality and survival of the generated VSMCs.
Dr Huiyu Zhou from the University of Leicester School of Computing and Mathematical Sciences is leading the research on artificial intelligence development by co-developing with other researchers a novel measurement technique, using spatial dispersion to quantify cytoskeletal organisation in F-actin staining and quantification process. This allowed the team to convert qualitative microscopy images into measurable data (intensity, orientation, density) to analyse cellular states and enables researchers to identify pathological, motile, or metabolic changes in cells
Dr Zhou said: “Our results suggest a promising treatment of CADASIL patients by PDE5 inhibitors to rescue VSMC function and prevent their degeneration seen in CADASIL small vessels. Our findings uncovered insights into the workings of posphodiesterase-5 and suggest a viable therapeutic strategy for small vessel disease and vascular dementia associated with NOTCH3, reinforcing the value of patient-specific iPSCs for disease modelling and potential drug discovery.
“The AI tool developed within this project helps quantify different classes of actin structure from stained confocal microscopy images, for studying actin related pathways involved in cell motility, and drug development.”
- ‘Selective vulnerability of cerebral vasculature to NOTCH3 variants in small vessel disease and rescue by phosphodiesterase-5 inhibitor’ is published in Science Advances, DOI: 10.1126/sciadv.aeb1134