Virtual Genetics Education Centre
Microbial genomes for higher education
Microbial genomes encompass all chromosomal and extra chromosomal genetic material. The study of genomes as an entity as opposed to individual genetic components is referred to as genomics.
Microbial genomes are widely variable and reflect the enormous diversity of bacteria, archaea and lower eukaryotes. Bacterial genomes usually consist of a single circular chromosome, but species with more than one chromosome (eg. Deinococcus radiodurans), linear chromosomes (eg. some Bacillis subtilis strains) and combinations of linear and circular chromosomes (eg. Agrobacterium tumefaciens) also exist.
The smallest microbial genome is that of Mycoplasma genitalium at 580,070 base pairs encoding 525 genes. Some of the largest genomes are those of the Cyanobacteria. Scytonema hofmanni's genome is 12,073,012 base pairs in size and has around 12,356 putative protein coding genes.
One major difference between the genomes of microorganisms and higher eukaryotes, is the presence of circular, extra-chromosomal DNA called plasmids. Plasmids can be transferred via horizontal DNA transfer from on cell of the same generation to another, mediating the rapid evolution of many different organisms.
The study of microbial genomes helps us to better understand the broader biology of bacteria, and how their genetic composition contributes to their tangible characteristics. The study of genomics is also important to infer the evolution of bacteria. Bacteria often evolve not just through small, single nucleotide level changes but through quantum evolutionary events. These include through the transfer of plasmids between species and also the transposition of large genetic elements within single cells. Understanding these processes allows us to determine the origins of bacteria and map the transfer of genes such as those conferring antibiotic resistance.
Genomics approaches allowed the authors to identify many different genes (particularly antibiotic resistance genes) that seem to have been acquired horizontally and which have driven the evolution of this superbug.
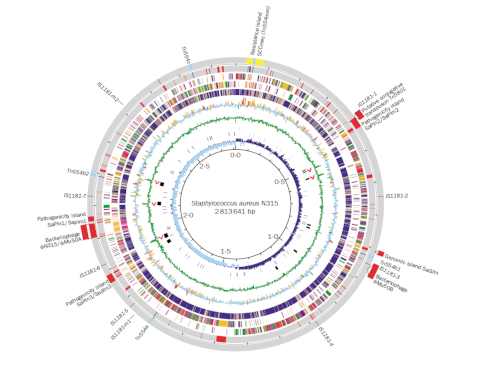
Studying microbial genomes
In the years preceding the development of full genome sequencing techniques we were restricted to studying microbial genomes in the lab using techniques such as PFGE.
Pulsed Field Gel Electrophoresis (PFGE)
The principle underlying PFGE is very similar to normal gel electrophoresis. PFGE however allows us to resolve far large, 'genome scale' pieces of DNA (greater than 20 kilobases in size). PFGE is still an important technique used to estimate the size of microbial genomes and in epidemiology studies.
DNA Sequencing
DNA sequencing has been an enormous advancement in the field of microbial genomics and indeed genetics as a whole, allowing us to amass vast amounts of genetic data form our organisms of choice. The first method for sequencing DNA was developed by Frederick Sanger and his group in 1977. Their method, termed Sanger Sequencing was a platform for innovation in the field of DNA sequencing, and we now have methods for sequencing entire bacterial genomes with relative ease. Whole genome sequencing produces immense amounts of data from which we can derive a catalogue of important information. From the need to analyse this data, the field of bioinformatics has flourished and become an integral part of genetics research.
DNA sequencing identifies and records every nucleotide, in order, that make up a piece of DNA.
Bioinformatics
Bioinformatics is the use of computing technologies to manage and analyse biological data. The huge amounts of genomic data produced through DNA sequencing can often be very confusing and difficult to analyse. Therefore bioinformatics is often required to derive useful information from this data. Using bioinformatics techniques, we can map the positions of genes, elicit their functions and infer evolutionary relationships. Together with experimental work we can use this information to learn more about how microbes ultimately survive and cause disease.