People
Professor Mark Carr
Professor of Structural Biology
School/Department: Molecular and Cell Biology, Department of
Email: mdc12@leicester.ac.uk
Research
Structure-based drug discovery
There is an urgent and unmet need for new drugs to treat major human diseases. Structure-based drug discovery, design and development (SBD3) has been developed as a major translational research focus by my group, with successful research partnerships built with UCB Biopharma, LifeArc (formerly MRC Technology) and a network of Cancer Research UK (CRUK) Centres across the UK. This work encompasses both small molecule and therapeutic antibody discovery programmes, with world-leading expertise developed in my group to use NMR-based methods to characterise complexes formed between potential therapeutic antibodies and target proteins. We are also developing innovative new approaches, such as antibody-assisted structure-based drug discovery.
The canonical Wnt signalling pathway has emerged as a key regulator of bone growth and remodelling, with activation of the pathway leading to increased bone density. Osteoporosis and related conditions occur as a consequence of the loss of bone integrity, and represent a major and growing medical challenge across the world. The activation of Wnt signalling is tightly regulated by a number of secreted proteins, including members of the Dickkopf family (Dkk-1, Dkk-2, Dkk-3 and Dkk-4) and sclerostin, which share the ability to interact with membrane-bound co-receptors for Wnt proteins (LRP5 and LRP6) leading to inhibition of Wnt signalling. These natural regulatory systems, together with the central role of Wnt signalling in bone homeostasis, highlight modulation of the regulation of Wnt signalling as an attractive target for the development of effective therapeutics to treat osteoporosis. Our collaboration with UCB Biopharma has resulted in determination of the first structure of sclerostin, as well as mapping of its interactions with functional partners and candidate therapeutic antibodies. In addition, we have recently determined the first structure of an N-terminal domain from a Dkk family protein (Dkk4).
Antibody-Assisted SBD3: The innovative use of antibodies as tools in SBD3 promises a number of important advances in knowledge-based drug discovery, including:
- The identification and characterisation of novel regulatory or allosteric sites on target proteins,
- The stabilisation and characterisation of rare, functionally important conformational states of target proteins, and
- The determination of structures for selected antibody-target protein complexes to guide the design of potential small molecule therapeutics
Importantly, antibody-assisted SBD3 has the potential to transform the ability to successfully target challenging protein therapeutic targets associated with major human diseases. We have recently established a pre-competitive Antibody-Assisted Structure-Based Drug Discovery Consortium with LifeArc and UCB Biopharma, which will jointly pioneer the development and application of antibody-assisted approaches.
Impact of SBD3 Programmes: The global impact of research in my group is exemplified by our mapping of the epitopes for a number of potential therapeutic antibodies, including the anti-TNF Cimzia (UCB Biopharma). Cimzia has now treated more than 98,000 patients in 62 countries with a range of major inflammatory diseases including rheumatoid arthritis, and Crohn’s disease, and has proved to be a highly effective therapeutic since its launch in 2009, which is reflected in sales of over 1.3 billion Euros in 2016. We have also made a major contribution to a UCB Biopharma programme targeting the treatment and prevention of osteoporosis by a therapeutic antibody (Romosozumab) against sclerostin, which is currently going through FDA approval and is expected to be transformative for osteoporosis treatment, as well as generate revenue of over a billion Euros per year. Similarly, our research partnership with LifeArc is also leading to new candidate therapeutics, with major contributions to at least two therapeutic antibody programmes that have now been licensed to industry for clinical development.
The global healthcare and massive economic impact of the SBD3 work in my group was recently recognised by the award of a Research Impact Award by the University for Best Economic Impact.
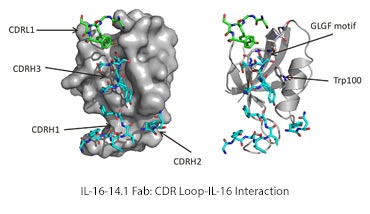
The figure shows molecular model of a human interleukin-16 shown as a spacefilling (left) and backbone only (right), bound to a fab fragment shown as stick representation, highlighting the binding groove on IL16
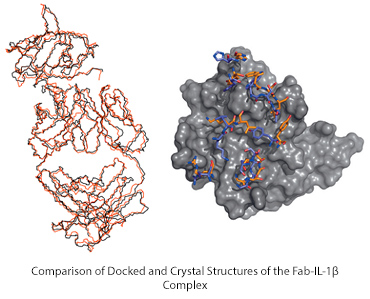
Overlay of two protein structures of the predicted and determined structures of interleukin 1beta and its antibody- backbones only to the left, surface representation to the right
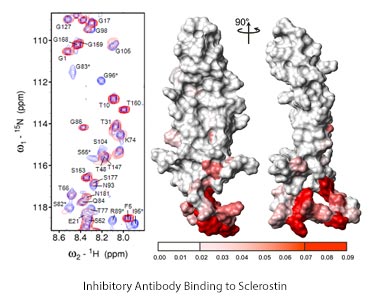
Left-overlaid 1H,15N HSQC NMR spectra of sclerostin, overlaying antibody- bound and unbound peaks. Right- mapping the shifted peaks onto the structure of sclerostin revealing the antibody binding site at the bottom of the protein as depicted.
Group members
Lorna Waters, Christine Prosser, Sarah Strong, Gareth Hall, Richard Cowan, Daniel Burschowsky, Chitra Seewooruthun, Yinan (Ellen) Fu, Jacqueline Kalms, Helena Wright, Roberta Baravalle, Matthew Powell-Grabaskey, Kirsty Ford, Jacqueline Kalms, Amaliawati Latiffi, Ra’eesah Shaikh, Tara Kang-Pettinger and Kayleigh Walker.
Publications
Key publications
- Chan DTY, Jenkinson L, Haynesa SW, Austin M, Diamandakis A, Burschowsky D, Seewooruthun C, Addyman A, Fiedler S, Ryman S, Whitehouse J, Slater LH, Gowans E, Shibata Y, Barnard M, Wilkinson RW, Vaughan TJ, Holt SV, Cerundolo V, Carr MD, Groves MAT (2020) Extensive sequence and structural evolution of Arginase2 inhibitory antibodies enabled by an unbiased approach to affinity maturation Proc. Natl. Acad Sci. (USA) 11.
- Schairer R et al (2020) 'Allosteric activation of MALT1 by its ubiquitin-binding Ig3 domain' Proceedings of the National Academy of Sciences DOI:10.1073/pnas.1912681117.
- Patel S, Barkell AM, Gupta D, Strong SL, Bruton S, Muskett FW, Addis PW, Renshaw PS, Slocombe PM, Doyle C, Clargo A, Taylor RJ, Prosser CE, Henry AJ, Robinson MK, Waters LC, Holdsworth G, Carr MD. (2018) Structural and functional analysis of Dickkopf 4 (Dkk4): New insights into Dkk evolution and regulation of Wnt signaling by Dkk and Kremen proteins. J Biol Chem. 293(31):12149-12166.
- Hall G, et al. (2016) 'Structure of a Potential Therapeutic Antibody Bound to Interleukin-16 (IL- 16): Mechanistic insights and new therapeutic opportunities.' Journal of Biological Chemistry, vol. 291. Pp. 16840–16848.
- Addis PW, et al. (2014) 'Conformational heterogeneity in antibody protein antigen recognition: implications for high affinity protein complex formation.' Journal of Biological Chemistry, vol. 289, no. Pp. 7200–7210.
- Cheng X, et al. (2013) 'Structure and interactions of the human programmed cell death 1 receptor.' Journal of Biological Chemistry, vol. 288. Pp. 11771–11785.
Press and media
Watch Mark's Bench to Business film.