People
Dr Amanda Chaplin
Lecturer in Structural Biology
School/Department: Molecular and Cell Biology, Department of
Email: ac853@leicester.ac.uk
Research
Structural architecture of DNA repair machinery
The major focus of our research is to understand the complex mechanisms underpinning DNA-repair using cryo-electron microscopy. Specifically, we work on proteins involved in the non-homologous end joining (NHEJ) pathway. NHEJ is crucial for cellular survival as it functions to repair DNA double-strand breaks (DSBs), which if not repaired correctly can lead to various diseases (most notably cancer).
My work has involved determining three-dimensional cryo-EM structures of the core proteins involved in NHEJ. Specifically, Ku70/80, DNA-PKcs, DNA-PK (the key enzyme) and supercomplexes including XLF, XRCC4 and Ligase IV. These structures have revealed the specific architecture of these elaborate complexes. We have also identified two novel dimeric forms of DNA-PK, allowing us to propose a new mechanism for NHEJ that utilises alternate oligomeric structural assemblies to respond to diverse DNA substrates.
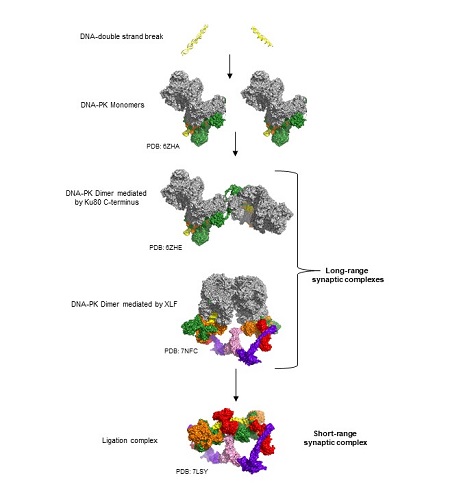
The diagram shows the steps involved in the non-homologous end joining pathway of DNA repair with protein structures. The panels show DNA doublestrand breaks, the assembly of long range synaptic complexes, followed by the ligation complex.
Our ultimate goal is to develop a complete picture of the mechanism of NHEJ. This will involve investigating the role of accessory proteins in NHEJ and how DNA-end configuration and packaging alter their recruitment and arrangement. There is now a greater understanding of the basics of NHEJ, but the adaptability of this system are governed by complexes it forms with accessory proteins. Using structural biology and biophysical techniques we will characterise different protein complexes involved at various stages of the mechanism. This work is in collaboration with Katheryn Meek (Michigan State University), Jean-Baptiste Charbonnier (Paris-Saclay) and Taiana Maia De Oliveira (AstraZeneca).
The NHEJ machinery is also involved in other cellular processes. One such example is during the innate immune response, where NHEJ proteins can detect viral DNA and trigger an antiviral response. However, many viruses have evolved methods to inhibit or modulate the host NHEJ proteins. We will use cryo-EM to probe interactions between viral proteins and the NHEJ machinery, with the ultimate aim of understanding how to develop novel anti-viral therapies. This work is in collaboration with Brian Ferguson (Cambridge).
Cryo-EM allows the possibility to visualise these structural assemblies, which could have significant impact on understanding disease progression and on human health. Both research areas allow for the development of drug-like molecules to target protein interaction sites, and ultimately combat cancer and viral infections.
Publications
Key publications
- Liang, S., Thomas S.E., Chaplin, A.K., Hardwick, S.W., Chirgadze, D.Y., Blundell, T.L. Structural insights into inhibitor regulation of the DNA repair protein DNA-PKcs Nature 601(7894):643-648.
- Chaplin, A.K., Hardwick, S.W., Liang, S., Kefala Stavridi, A., Hnizda, A., Cooper, L.R., De Oliveira, T.M., Chirgadze, D.Y., and Blundell, T.L. 2020. Dimers of DNA-PK create a stage for DNA double-strand break repair. Nature Structural and Molecular Biology, 28, 13-19.
- Chaplin, A.K, Hardwick, S.W., Kefala Stavridi, A., Buehl, C., Goff, N. J., Ropars, V., Liang, S., De Oliveira, T.M., Chirgadze, D.Y., Meek, K., Charbonnier, J. B., Blundell, T.L. 2021. Cryo-EM of NHEJ supercomplexes provides insights into DNA repair. Molecular Cell. 81. 3400-3409.